When referencing this material, please cite:
Please also include the standard citation for PhysioNet:
The Cyclic Alternating Pattern (CAP) of EEG activity during sleep
The Cyclic Alternating Pattern (CAP) is a periodic EEG activity occurring during NREM sleep. It is characterized by cyclic sequences of cerebral activation (phase A) followed by periods of deactivation (phase B) which separate two successive phase A periods with an interval <1 min. A phase A period and the following phase B period define a CAP cycle, and at least two CAP cycles are required to form a CAP sequence.
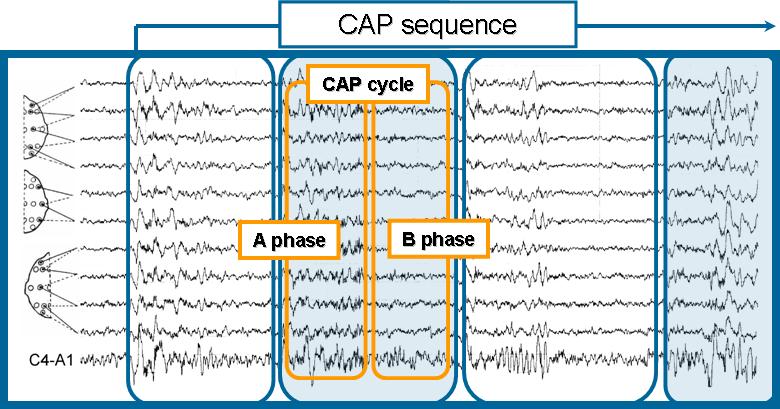
An example of cyclic alternating pattern (CAP) in sleep stage 2. A CAP cycle is defined as a phase A period followed by a phase B period lasting a minute or less. Two or more adjacent CAP cycles define a CAP sequence.
Phase A periods are subdivided into three subtypes:
- Subtype A1: synchronized events with low impact on autonomic and somatomotor activities;
- Subtype A2: mixed synchronized–desynchronized EEG events with an intermediate influence on the autonomic and somatomotor activities; and
- Subtype A3: predominantly desynchronized EEG events with heavy effects on the autonomic and somatomotor activities.
A detailed description and definition of CAP is reported in [1]. For a complete review of the history and significance of CAP, see [2].
Despite being a physiological phenomenon, CAP is also a marker of sleep instability and can be correlated with several sleep-related pathologies. In fact, on one hand CAP can reflect a reaction of the sleeping brain to any endogenous or exogenous disturbance; on the other hand, the A phase of CAP has been interpreted as a kind of gate through which pathological events more easily occur [3]. Increased amounts of CAP are often observed in sleep-disordered breathing (SDB), as well as in insomnia [4], sleep movement disorders (periodic leg movements, or PLM [5], restless leg syndrome (RLS) [6], parasomnias like REM behavior disorder (RBD), and epileptic diseases such as nocturnal frontal lobe epilepsy (NFLE) [7]. Pathological amounts of CAP can also be found in hypersomnias of central origin such as narcolepsy [8].
The ratio between NREM CAP sleep and total NREM sleep (CAP-rate), and the different distributions of CAP A phases with respect to sleep stages can be measured in sleep centers to characterize such pathologies. These indexes can be valuable as measures of sleep quality, but the time needed to determine them has been an obstacle to their adoption in routine clinical practice, since visual scoring of CAP phases A for an entire night of sleep recording requires expertise and lengthy, attentive analysis. The development of software capable of increasingly accurate and efficient automated analysis of CAP is beginning to result in wider clinical use of CAP indexes for diagnosis of sleep pathologies and assessment of the efficacy of therapeutic interventions.
The CAP Sleep Database
The CAP Sleep Database is a collection of 108 polysomnographic recordings registered at the Sleep Disorders Center of the Ospedale Maggiore of Parma, Italy. The waveforms (contained in the .edf files of the database) include at least 3 EEG channels (F3 or F4, C3 or C4 and O1 or O2, referred to A1 or A2), EOG (2 channels), EMG of the submentalis muscle, bilateral anterior tibial EMG, respiration signals (airflow, abdominal and thoracic effort and SaO2) and EKG. Additional traces include EEG bipolar traces, according to the 10-20 international system (Fp1-F3, F3-C3, C3-P3, P3-O1 and/or Fp2-F4, F4-C4, C4-P4, P4-O2).
The 16 healthy subjects included in the study did not present any neurological disorders and were free of drugs affecting the central nervous system. The 92 pathological recordings include 40 recordings of patients diagnosed with NFLE, 22 affected by RBD, 10 with PLM, 9 insomniac, 5 narcoleptic, 4 affected by SDB and 2 by bruxism. The age and gender of the subjects are reported in a spreadsheet (gender-age.xlsx).
The recordings are named according to the subjects' pathology:
| n1-n16 | No pathology (controls) |
| brux1-brux2 | Bruxism |
| ins1-ins9 | Insomnia |
| narco1-narco5 | Narcolepsy |
| nfle1-nfle40 | Nocturnal frontal lobe epilepsy |
| plm1-plm10 | Periodic leg movements |
| rbd1-rbd22 | REM behavior disorder |
| sdb1-sdb4 | Sleep-disordered breathing |
Annotations
Expert neurologists trained at the Sleep Center provided the scoring of the sleep macrostructure, according to the Rechtschaffen & Kales rules [9], while the CAP was detected in agreement with Terzano’s reference atlas of rules [10]. All the signals were visualized and the scorings were performed using REMlogic™ software (Embla).
The scores for each recording are provided as .txt files in REMlogic report format, and also as .st files in PhysioBank-compatible format. The .txt score files have the following fields:
- Sleep stage (W=wake, S1-S4=sleep stages, R=REM, MT=body movements)
- Body position (Left, Right, Prone, or Supine; not recorded in some subjects)
- Time of day [hh:mm:ss]
- Event (either a sleep stage (SLEEP-S0..S4, REM, MT), or a phase A of CAP)
- Duration (in seconds)
- Location (the signal(s) in which the event can be observed)
A Matlab script is provided for reading the .txt score files easily. By launching ScoringReader.m, an array containing the scoring of the macrostructure, and three arrays containing the starting time of the CAP A phases, their duration and their subtype, are generated. The hypnogram is reported as 0 for wake, 1 to 4 for the sleep stages, according to R&K rules, and 5 for REM. The Matlab function CAP.m computes, starting from these variables, the CAP time and CAP rate according to Terzano’s rules.
The .st score files can be read using the PhysioBank ATM and other software that reads PhysioBank-compatible annotation files. These files contain the same information as the .txt files; times are encoded as for other annotation files, and each annotation's aux string contains the associated event, duration, sleep stage, location, and (where available) body position.
Applications for this database
This database is intended to provide a useful number of carefully annotated examples of CAP in a representative variety of pathophysiologic contexts, for development and evaluation of automated CAP analyzers, as well as to support basic studies of the dynamics of CAP.
Several efforts have been made to develop a reliable automatic CAP-scoring algorithm [11]. Most of these methods rely on the extraction of spectral features from the EEG and on the application of machine-learning algorithms. Although all these methods achieve good results, none of them is yet applied to clinical practice since they either require some amount of clinician intervention or do not achieve a sufficient accuracy in the classification to support the clinical diagnosis. Some recordings from this database have been used for studies on features related to CAP phase A [12-16] and for training automatic CAP detection algorithms [17].
Acknowledgements
The CAP Sleep Database was contributed to PhysioNet by Professor Mario Giovanni Terzano, MD and Professor Liborio Parrino, MD. The team of sleep experts of the Sleep Disorders Center of the Ospedale Maggiore of Parma, Italy, who performed the scoring includes Andrea Grassi, MD, Fernando De Paolis, MD, Giulia Milioli, MD, Silvia Riccardi, MD, Nicoletta Azzi, BSc, and Valentina Rosso, BSc. The BIOSIP group at the Bioengineering Department of Politecnico di Milano, Italy contributed to the organization of the database and implemented the Matlab scripts.
PhysioNet also gratefully acknowledges the contributions of BIOSIP group member Sara Mariani, MSc, who coordinated the preparation of the database for public release and its documentation and related software as a PhysioNetWorks project during her visit to MIT.
Bibliography
- MG Terzano, D Mancia, MR Salati, G Costani, A Decembrino, L Parrino. The cyclic alternating pattern as a physiologic component of normal NREM sleep. Sleep 1985;8(2):137-145.
- L Parrino, R Ferri, O Bruni, M. G. Terzano. Cyclic alternating pattern (CAP): The marker of sleep instability. Sleep Med Rev 2012 Feb;16(1):27-45
- L Parrino, P Halasz, CA Tassinari, MG Terzano.C CAP, epilepsy and motor events during sleep: the unifying role of arousal. Sleep Med Rev 2006 Aug;10(4):267-285.
- MG Terzano, L Parrino, MC Spaggiari, V Palomba, M Rossi, A Smerieri. CAP variables and arousals as sleep electroencephalogram markers for primary insomnia. Clin Neurophysiol 2003 Sep;114(9):1715-1723.
- L Parrino, M Boselli, GP Buccino, MC Spaggiari, G Di Giovanni, MG Terzano. The cyclic alternating pattern plays a gate-control on periodic limb movements during non-rapid eye movement sleep. J Clin Neurophysiol 1996 Jul;13(4):314-323.
- W Hening. The clinical neurophysiology of the restless legs syndrome and periodic limb movements. Part I: diagnosis, assessment, and characterization. Clin Neurophysiol 2004 Sep;115(9):1965-1974.
- T Kato, JY Montplaisir, F Guitard, BJ Sessle, JP Lund, GJ Lavigne. Evidence that experimentally induced sleep bruxism is a consequence of transient arousal. J Dent Res 2003 Apr;82(4):284-288.
- M Zucconi, L Ferini-Strambi. NREM parasomnias: arousal disorders and differentiation from nocturnal frontal lobe epilepsy. Clin Neurophysiol 2000 Sep;111 Suppl 2:S129-S135.
- R Poryazova, E Werth, L Parrino, MG Terzano, CL Bassetti. Cyclic alternating pattern in narcolepsy patients and healthy controls after partial and total sleep deprivation. Clin Neurophysiol 2011 Sep;122(9):1788-1793.
- A Rechtscahffen, A. Kales. A manual of standardized terminology, techniques and scoring system for sleep stages in human subjects. National Institutes of Health, Neurological Information Network, Publication 204, 1968.
- MG Terzano, L Parrino, A Sherieri, R Chervin, S Chokroverty, C Guilleminault, M Hirshkowitz, M Mahowald, H Moldofsky, A Rosa, R Thomas, A Walters. Atlas, rules, and recording techniques for the scoring of cyclic alternating pattern (CAP) in human sleep. Sleep Med 2001 Nov;2(6):537-553.
- U Barcaro, E Bonanni, M Maestri, L Murri, L Parrino, MG Terzano. A general automatic method for the analysis of NREM sleep microstructure. Sleep Med 2004 Nov;5(6):567-576.
- R Largo, C Munteanu, A Rosa. CAP event detection by wavelets and GA tuning. Proc 2005 IEEE International Workshop on Intelligent Signal Processing 2005, pp. 44-48.
- R Ferri, O Bruni, S Miano, A Smerieri, K Spruyt, MG Terzano. Inter-rater reliability of sleep cyclic alternating pattern (CAP) scoring and validation of a new computer-assisted CAP scoring method. Clin Neurophysiol 2005 Mar;116(3):696-707.
- S Mariani, AM Bianchi, E Manfredini, V Rosso, MO Mendez, L Parrino, M Matteucci, A Grassi, S Cerutti, MG Terzano. Automatic detection of A phases of the cyclic alternating pattern during sleep. Proc IEEE Eng Med Biol Soc 2010;5085-5088.
- C Navona, U Barcaro, E Bonanni, F Di Martino, M Maestri, L Murri. An automatic method for the recognition and classification of the A-phases of the cyclic alternating pattern. Clin Neurophysiol 2002 Nov;113(11):1826-1831.
- S Mariani, E Manfredini, V Rosso, MO Mendez, AM Bianchi, M Matteucci, MG Terzano, S Cerutti, L Parrino. Characterization of A phases during the Cyclic Alternating Pattern of sleep. Clin Neurophysiol 2011 Oct;122(10):2016-2024.
Database files
Name Last modified Size Description
Parent Directory -
gender-age.xlsx 2011-08-11 19:35 11K
CAP.m 2011-08-12 13:39 1.7K
ScoringReader.m 2011-08-19 17:30 4.2K
CAP-sequence.jpg 2012-07-24 10:37 81K
RECORDS 2012-07-24 18:49 1.0K list of record names
brux1.edf.st 2012-07-25 14:50 12K
brux2.edf.st 2012-07-25 14:50 47K
ins1.edf.st 2012-07-25 14:50 40K
ins2.edf.st 2012-07-25 14:50 64K
ins3.edf.st 2012-07-25 14:50 36K
ins4.edf.st 2012-07-25 14:50 34K
ins5.edf.st 2012-07-25 14:50 68K
ins6.edf.st 2012-07-25 14:50 43K
ins7.edf.st 2012-07-25 14:50 61K
ins8.edf.st 2012-07-25 14:50 39K
ins9.edf.st 2012-07-25 14:50 42K
n10.edf.st 2012-07-25 14:50 35K
n11.edf.st 2012-07-25 14:50 44K
n12.edf.st 2012-07-25 14:50 37K
n13.edf.st 2012-07-25 14:50 45K
n14.edf.st 2012-07-25 14:50 43K
n1.edf.st 2012-07-25 14:50 52K
n15.edf.st 2012-07-25 14:50 42K
n16.edf.st 2012-07-25 14:50 43K
n2.edf.st 2012-07-25 14:50 42K
n3.edf.st 2012-07-25 14:50 42K
n4.edf.st 2012-07-25 14:50 51K
n5.edf.st 2012-07-25 14:50 49K
n6.edf.st 2012-07-25 14:50 45K
n7.edf.st 2012-07-25 14:50 42K
n8.edf.st 2012-07-25 14:50 49K
n9.edf.st 2012-07-25 14:50 39K
narco1.edf.st 2012-07-25 14:50 41K
narco2.edf.st 2012-07-25 14:50 12K
narco3.edf.st 2012-07-25 14:50 64K
narco4.edf.st 2012-07-25 14:50 48K
narco5.edf.st 2012-07-25 14:50 32K
nfle10.edf.st 2012-07-25 14:50 46K
nfle11.edf.st 2012-07-25 14:50 49K
nfle12.edf.st 2012-07-25 14:50 51K
nfle13.edf.st 2012-07-25 14:50 59K
nfle14.edf.st 2012-07-25 14:50 44K
nfle15.edf.st 2012-07-25 14:50 50K
nfle16.edf.st 2012-07-25 14:50 52K
nfle1.edf.st 2012-07-25 14:50 49K
nfle17.edf.st 2012-07-25 14:50 47K
nfle18.edf.st 2012-07-25 14:50 52K
nfle19.edf.st 2012-07-25 14:50 49K
nfle2.edf.st 2012-07-25 14:50 48K
nfle20.edf.st 2012-07-25 14:50 45K
nfle21.edf.st 2012-07-25 14:50 64K
nfle22.edf.st 2012-07-25 14:50 54K
nfle23.edf.st 2012-07-25 14:50 47K
nfle24.edf.st 2012-07-25 14:50 48K
nfle25.edf.st 2012-07-25 14:50 50K
nfle26.edf.st 2012-07-25 14:50 44K
nfle27.edf.st 2012-07-25 14:50 34K
nfle28.edf.st 2012-07-25 14:50 51K
nfle29.edf.st 2012-07-25 14:50 48K
nfle30.edf.st 2012-07-25 14:50 46K
nfle31.edf.st 2012-07-25 14:50 40K
nfle32.edf.st 2012-07-25 14:50 52K
nfle33.edf.st 2012-07-25 14:50 51K
nfle34.edf.st 2012-07-25 14:50 47K
nfle35.edf.st 2012-07-25 14:50 40K
nfle36.edf.st 2012-07-25 14:50 47K
nfle37.edf.st 2012-07-25 14:50 46K
nfle38.edf.st 2012-07-25 14:50 49K
nfle3.edf.st 2012-07-25 14:50 50K
nfle39.edf.st 2012-07-25 14:50 46K
nfle4.edf.st 2012-07-25 14:50 53K
nfle40.edf.st 2012-07-25 14:50 58K
nfle5.edf.st 2012-07-25 14:50 52K
nfle6.edf.st 2012-07-25 14:50 35K
nfle7.edf.st 2012-07-25 14:50 63K
nfle8.edf.st 2012-07-25 14:50 45K
nfle9.edf.st 2012-07-25 14:50 54K
plm1.edf.st 2012-07-25 14:50 55K
plm10.edf.st 2012-07-25 14:50 46K
plm2.edf.st 2012-07-25 14:50 40K
plm3.edf.st 2012-07-25 14:50 49K
plm4.edf.st 2012-07-25 14:50 41K
plm5.edf.st 2012-07-25 14:50 39K
plm6.edf.st 2012-07-25 14:50 33K
plm7.edf.st 2012-07-25 14:50 44K
plm8.edf.st 2012-07-25 14:50 37K
plm9.edf.st 2012-07-25 14:50 42K
rbd10.edf.st 2012-07-25 14:50 34K
rbd11.edf.st 2012-07-25 14:50 33K
rbd12.edf.st 2012-07-25 14:50 45K
rbd13.edf.st 2012-07-25 14:50 57K
rbd1.edf.st 2012-07-25 14:50 29K
rbd14.edf.st 2012-07-25 14:50 49K
rbd15.edf.st 2012-07-25 14:50 39K
rbd16.edf.st 2012-07-25 14:50 48K
rbd17.edf.st 2012-07-25 14:50 50K
rbd18.edf.st 2012-07-25 14:50 40K
rbd19.edf.st 2012-07-25 14:50 46K
rbd2.edf.st 2012-07-25 14:50 60K
rbd20.edf.st 2012-07-25 14:50 35K
rbd21.edf.st 2012-07-25 14:50 53K
rbd22.edf.st 2012-07-25 14:50 48K
rbd3.edf.st 2012-07-25 14:50 56K
rbd4.edf.st 2012-07-25 14:50 57K
rbd5.edf.st 2012-07-25 14:50 54K
rbd6.edf.st 2012-07-25 14:50 30K
rbd7.edf.st 2012-07-25 14:50 44K
rbd8.edf.st 2012-07-25 14:50 52K
rbd9.edf.st 2012-07-25 14:50 45K
sdb1.edf.st 2012-07-25 14:50 32K
sdb2.edf.st 2012-07-25 14:50 48K
sdb3.edf.st 2012-07-25 14:50 37K
sdb4.edf.st 2012-07-25 14:50 59K
ANNOTATORS 2012-07-25 14:56 43 list of annotators
DOI 2015-09-21 13:00 20
sdb4.edf 2018-11-09 14:55 206M
brux1.edf 2018-11-09 14:56 221M
brux2.edf 2018-11-09 14:56 252M
ins1.edf 2018-11-09 14:57 253M
ins2.edf 2018-11-09 14:58 466M
ins3.edf 2018-11-09 14:58 166M
ins4.edf 2018-11-09 14:58 270M
ins5.edf 2018-11-09 14:59 378M
ins6.edf 2018-11-09 15:00 448M
ins7.edf 2018-11-09 15:00 326M
ins8.edf 2018-11-09 15:01 418M
ins9.edf 2018-11-09 15:02 520M
n1.edf 2018-11-09 15:02 473M
n2.edf 2018-11-09 15:03 356M
n3.edf 2018-11-09 15:04 525M
n5.edf 2018-11-09 15:05 499M
n6.edf 2018-11-09 15:05 62M
n7.edf 2018-11-09 15:05 58M
n9.edf 2018-11-09 15:06 62M
n10.edf 2018-11-09 15:06 452M
n11.edf 2018-11-09 15:07 494M
n12.edf 2018-11-09 15:07 51M
n13.edf 2018-11-09 15:08 118M
n14.edf 2018-11-09 15:08 124M
n16.edf 2018-11-09 15:08 29M
narco1.edf 2018-11-09 15:09 318M
narco2.edf 2018-11-09 15:09 257M
narco3.edf 2018-11-09 15:10 416M
narco4.edf 2018-11-09 15:11 518M
narco5.edf 2018-11-09 15:11 191M
nfle1.edf 2018-11-09 15:12 510M
nfle2.edf 2018-11-09 15:12 491M
nfle3.edf 2018-11-09 15:13 572M
nfle4.edf 2018-11-09 15:14 551M
nfle5.edf 2018-11-09 15:15 518M
nfle6.edf 2018-11-09 15:15 153M
nfle7.edf 2018-11-09 15:16 534M
nfle8.edf 2018-11-09 15:17 232M
nfle9.edf 2018-11-09 15:18 601M
nfle10.edf 2018-11-09 15:18 158M
nfle11.edf 2018-11-09 15:18 165M
nfle12.edf 2018-11-09 15:19 547M
nfle13.edf 2018-11-09 15:20 514M
nfle14.edf 2018-11-09 15:21 482M
nfle15.edf 2018-11-09 15:22 544M
nfle16.edf 2018-11-09 15:22 525M
nfle17.edf 2018-11-09 15:26 457M
nfle18.edf 2018-11-09 15:27 542M
nfle19.edf 2018-11-09 15:28 146M
nfle20.edf 2018-11-09 15:28 183M
nfle21.edf 2018-11-09 15:29 542M
nfle22.edf 2018-11-09 15:30 531M
nfle23.edf 2018-11-09 15:30 169M
nfle24.edf 2018-11-09 15:31 482M
nfle25.edf 2018-11-09 15:31 226M
nfle26.edf 2018-11-09 15:31 157M
nfle27.edf 2018-11-09 15:32 556M
nfle28.edf 2018-11-09 15:33 567M
nfle29.edf 2018-11-09 15:34 477M
nfle30.edf 2018-11-09 15:35 545M
nfle31.edf 2018-11-09 15:35 178M
nfle32.edf 2018-11-09 15:36 479M
nfle33.edf 2018-11-09 15:37 739M
nfle34.edf 2018-11-09 15:38 526M
nfle35.edf 2018-11-09 15:39 471M
nfle36.edf 2018-11-09 15:40 520M
nfle37.edf 2018-11-09 15:41 480M
nfle38.edf 2018-11-09 15:41 529M
nfle39.edf 2018-11-09 15:42 471M
nfle40.edf 2018-11-09 15:43 563M
plm1.edf 2018-11-09 15:44 276M
plm2.edf 2018-11-09 15:45 357M
plm3.edf 2018-11-09 15:45 382M
plm4.edf 2018-11-09 15:46 326M
plm5.edf 2018-11-09 15:46 428M
plm6.edf 2018-11-09 15:47 500M
plm7.edf 2018-11-09 15:48 293M
plm8.edf 2018-11-09 15:48 232M
plm9.edf 2018-11-09 15:49 491M
plm10.edf 2018-11-09 15:49 360M
rbd1.edf 2018-11-09 15:50 303M
rbd2.edf 2018-11-09 15:51 542M
rbd3.edf 2018-11-09 15:52 572M
rbd4.edf 2018-11-09 15:53 523M
rbd5.edf 2018-11-09 15:54 484M
rbd6.edf 2018-11-09 15:54 527M
rbd7.edf 2018-11-09 15:55 444M
rbd8.edf 2018-11-09 15:56 449M
rbd9.edf 2018-11-09 15:57 485M
rbd10.edf 2018-11-09 15:57 358M
rbd11.edf 2018-11-09 15:58 358M
rbd12.edf 2018-11-09 15:59 456M
rbd13.edf 2018-11-09 15:59 489M
rbd14.edf 2018-11-09 16:00 339M
rbd15.edf 2018-11-09 16:01 485M
rbd16.edf 2018-11-09 16:02 471M
rbd17.edf 2018-11-09 16:02 367M
rbd18.edf 2018-11-09 16:03 516M
rbd19.edf 2018-11-09 16:04 522M
rbd20.edf 2018-11-09 16:04 297M
rbd21.edf 2018-11-09 16:05 460M
rbd22.edf 2018-11-09 16:06 484M
sdb1.edf 2018-11-09 16:06 139M
sdb2.edf 2018-11-09 16:06 143M
sdb3.edf 2018-11-09 16:07 308M
n15.edf 2018-11-14 16:39 124M
n4.edf 2018-11-14 16:40 123M
n8.edf 2018-11-14 16:40 88M
brux1.txt 2018-11-16 13:51 72K
brux2.txt 2018-11-16 13:51 74K
ins1.txt 2018-11-16 13:51 42K
ins2.txt 2018-11-16 13:51 65K
ins3.txt 2018-11-16 13:51 38K
ins4.txt 2018-11-16 13:51 35K
ins5.txt 2018-11-16 13:51 70K
ins6.txt 2018-11-16 13:51 60K
ins7.txt 2018-11-16 13:51 63K
ins8.txt 2018-11-16 13:51 61K
ins9.txt 2018-11-16 13:51 65K
n1.txt 2018-11-16 13:51 82K
n10.txt 2018-11-16 13:51 55K
n11.txt 2018-11-16 13:51 70K
n12.txt 2018-11-16 13:51 60K
n13.txt 2018-11-16 13:51 71K
n14.txt 2018-11-16 13:51 68K
n15.txt 2018-11-16 13:51 69K
n16.txt 2018-11-16 13:51 73K
n2.txt 2018-11-16 13:51 44K
n3.txt 2018-11-16 13:51 44K
n4.txt 2018-11-16 13:51 53K
n5.txt 2018-11-16 13:51 77K
n6.txt 2018-11-16 13:51 47K
n7.txt 2018-11-16 13:51 68K
n8.txt 2018-11-16 13:51 51K
n9.txt 2018-11-16 13:51 63K
narco1.txt 2018-11-16 13:51 43K
narco2.txt 2018-11-16 13:51 65K
narco3.txt 2018-11-16 13:51 94K
narco4.txt 2018-11-16 13:51 75K
narco5.txt 2018-11-16 13:51 45K
nfle1.txt 2018-11-16 13:51 77K
nfle10.txt 2018-11-16 13:51 72K
nfle11.txt 2018-11-16 13:51 78K
nfle12.txt 2018-11-16 13:51 53K
nfle13.txt 2018-11-16 13:51 93K
nfle14.txt 2018-11-16 13:51 69K
nfle15.txt 2018-11-16 13:51 80K
nfle16.txt 2018-11-16 13:51 81K
nfle17.txt 2018-11-16 13:51 73K
nfle18.txt 2018-11-16 13:51 81K
nfle19.txt 2018-11-16 13:51 77K
nfle2.txt 2018-11-16 13:51 51K
nfle20.txt 2018-11-16 13:51 72K
nfle21.txt 2018-11-16 13:51 67K
nfle22.txt 2018-11-16 13:51 56K
nfle23.txt 2018-11-16 13:51 75K
nfle24.txt 2018-11-16 13:51 76K
nfle25.txt 2018-11-16 13:51 80K
nfle26.txt 2018-11-16 13:51 68K
nfle27.txt 2018-11-16 13:51 59K
nfle28.txt 2018-11-16 13:51 80K
nfle29.txt 2018-11-16 13:51 50K
nfle3.txt 2018-11-16 13:51 78K
nfle30.txt 2018-11-16 13:51 72K
nfle31.txt 2018-11-16 13:51 64K
nfle32.txt 2018-11-16 13:51 54K
nfle33.txt 2018-11-16 13:51 79K
nfle34.txt 2018-11-16 13:51 74K
nfle35.txt 2018-11-16 13:51 42K
nfle36.txt 2018-11-16 13:51 73K
nfle37.txt 2018-11-16 13:51 74K
nfle38.txt 2018-11-16 13:51 77K
nfle39.txt 2018-11-16 13:51 72K
nfle4.txt 2018-11-16 13:52 84K
nfle40.txt 2018-11-16 13:52 92K
nfle5.txt 2018-11-16 13:52 54K
nfle6.txt 2018-11-16 13:52 55K
nfle7.txt 2018-11-16 13:52 100K
nfle8.txt 2018-11-16 13:52 72K
nfle9.txt 2018-11-16 13:52 56K
plm1.txt 2018-11-16 13:52 87K
plm10.txt 2018-11-16 13:52 72K
plm2.txt 2018-11-16 13:52 63K
plm3.txt 2018-11-16 13:52 78K
plm4.txt 2018-11-16 13:52 43K
plm5.txt 2018-11-16 13:52 41K
plm6.txt 2018-11-16 13:52 51K
plm7.txt 2018-11-16 13:52 69K
plm8.txt 2018-11-16 13:52 39K
plm9.txt 2018-11-16 13:52 66K
rbd1.txt 2018-11-16 13:52 46K
rbd10.txt 2018-11-16 13:52 52K
rbd11.txt 2018-11-16 13:52 35K
rbd12.txt 2018-11-16 13:52 47K
rbd13.txt 2018-11-16 13:52 59K
rbd14.txt 2018-11-16 13:52 76K
rbd15.txt 2018-11-16 13:52 60K
rbd16.txt 2018-11-16 13:52 76K
rbd17.txt 2018-11-16 13:52 78K
rbd18.txt 2018-11-16 13:52 64K
rbd19.txt 2018-11-16 13:52 73K
rbd2.txt 2018-11-16 13:52 94K
rbd20.txt 2018-11-16 13:52 55K
rbd21.txt 2018-11-16 13:52 55K
rbd22.txt 2018-11-16 13:52 75K
rbd3.txt 2018-11-16 13:52 58K
rbd4.txt 2018-11-16 13:52 59K
rbd5.txt 2018-11-16 13:52 56K
rbd6.txt 2018-11-16 13:52 31K
rbd7.txt 2018-11-16 13:52 69K
rbd8.txt 2018-11-16 13:52 82K
rbd9.txt 2018-11-16 13:52 71K
sdb1.txt 2018-11-16 13:52 51K
sdb2.txt 2018-11-16 13:52 76K
sdb3.txt 2018-11-16 13:52 39K
sdb4.txt 2018-11-16 13:52 95K
MD5SUMS 2018-11-16 14:10 14K
SHA1SUMS 2018-11-16 14:10 17K
SHA256SUMS 2018-11-16 14:10 25K
|
If you would like help understanding, using, or downloading content, please see our Frequently Asked Questions. If you have any comments, feedback, or particular questions regarding this page, please send them to the webmaster. Comments and issues can also be raised on PhysioNet's GitHub page. Updated Friday, 28 October 2016 at 16:58 EDT |
PhysioNet is supported by the National Institute of General Medical Sciences (NIGMS) and the National Institute of Biomedical Imaging and Bioengineering (NIBIB) under NIH grant number 2R01GM104987-09.
|