This database is used and described in
Please cite this publication when referencing this material, and also include the standard citation for PhysioNet:
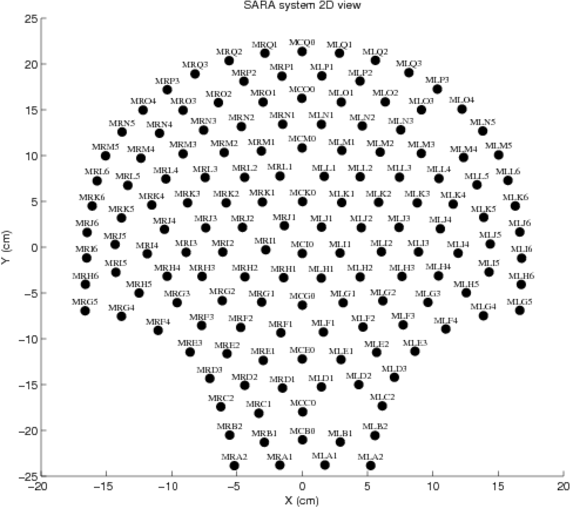
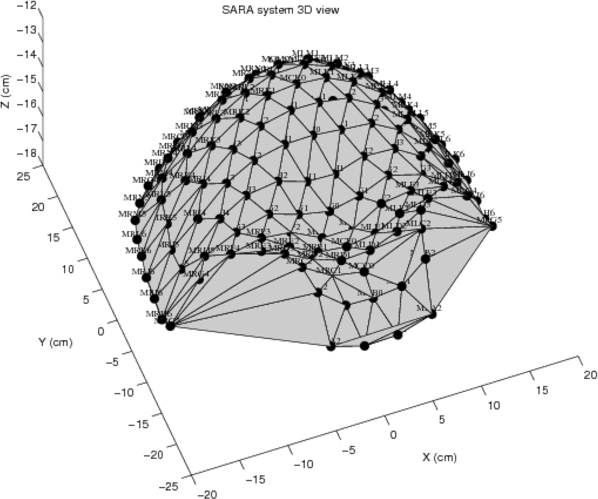
This database contains uterine magnetomyographic (MMG) signals recorded using the 151 channel SARA (SQUID Array for Reproductive Assessment) system installed at UAMS, Little Rock, USA. Recordings are taken from 25 subjects who presented themselves in the Triage unit of Labor and Delivery and were undergoing monitoring and evaluation of labor. All subjects were asked to lean forward and sit comfortably with the sensor array covering their pregnant abdomens. The duration of a recording was typically around 20 minutes.
To obtain the final MMG signals, the original 250Hz data was downsampled to 32 Hz, bandpass filtered (0.1-1 Hz) to attenuate the maternal and fetal cardiac signals, and notch filtered (0.25-0.35 Hz) to suppress the prominent maternal breathing around 0.33 Hz. Furthermore, segments with maternal movement were excluded. Each final recording in this database contains 147 to 148 channels and lasts between 10 and 20 minutes.
The data was preprocessed in MATLAB and converted to WFDB format using the mat2wfdb function from the PhysioToolkit software. File names in the database are of the form ###_*GA, where ### is the ID of the participant, and *GA refers to gestational age in weeks and days (ie. 38w0d). Header files contain channel labels and the following clinical information:
- Cervical dilation (dilation cm)/(effacement %)/(station)
- Days to delivery after SARA recording.
- Race
- BMI
- Gestation Period (weeks+days)
The sensorpositions record contains x, y, and z coordinates of the SARA array's 149 different channels present in the database. Refer to SARAarray.eps or SARAarray.tif for diagrams of the apparatus.
Relevant Publications
Name Last modified Size Description
Parent Directory -
202_38w0d.dat 2016-03-02 19:26 9.2M digitized signal(s)
202_38w0d.hea 2016-03-02 19:27 8.2K header file
203_37w3d.dat 2016-03-02 19:26 8.1M digitized signal(s)
203_37w3d.hea 2016-03-02 19:27 8.3K header file
204_37w6d.dat 2016-03-02 19:26 10M digitized signal(s)
204_37w6d.hea 2016-03-02 19:27 8.3K header file
205_40w1d.dat 2016-03-02 19:26 7.6M digitized signal(s)
205_40w1d.hea 2016-03-02 19:27 8.3K header file
207_38w2d.dat 2016-03-02 19:26 9.2M digitized signal(s)
207_38w2d.hea 2016-03-02 19:27 8.3K header file
209_37w3d.dat 2016-03-02 19:26 9.8M digitized signal(s)
209_37w3d.hea 2016-03-02 19:27 8.3K header file
210_37w3d.dat 2016-03-02 19:26 9.8M digitized signal(s)
210_37w3d.hea 2016-03-02 19:27 8.3K header file
211_37w2d.dat 2016-03-02 19:26 9.2M digitized signal(s)
211_37w2d.hea 2016-03-02 19:27 8.3K header file
212_39w0d.dat 2016-03-02 19:26 7.6M digitized signal(s)
212_39w0d.hea 2016-03-02 19:27 8.3K header file
213_37w4d.dat 2016-03-02 19:26 7.6M digitized signal(s)
213_37w4d.hea 2016-03-02 19:27 8.3K header file
214_37w3d.dat 2016-03-02 19:26 7.0M digitized signal(s)
214_37w3d.hea 2016-03-02 19:27 8.2K header file
218_38w0d.dat 2016-03-02 19:26 7.6M digitized signal(s)
218_38w0d.hea 2016-03-02 19:27 8.3K header file
221_37w0d.dat 2016-03-02 19:26 9.2M digitized signal(s)
221_37w0d.hea 2016-03-02 19:27 8.2K header file
222_38w0d.dat 2016-03-02 19:26 9.2M digitized signal(s)
222_38w0d.hea 2016-03-02 19:27 8.3K header file
224_39w0d.dat 2016-03-02 19:26 9.5M digitized signal(s)
224_39w0d.hea 2016-03-02 19:27 8.2K header file
225_37w6d.dat 2016-03-02 19:26 6.2M digitized signal(s)
225_37w6d.hea 2016-03-02 19:27 8.2K header file
226_38w0d.dat 2016-03-02 19:26 8.4M digitized signal(s)
226_38w0d.hea 2016-03-02 19:27 8.3K header file
227_40w3d.dat 2016-03-02 19:26 9.5M digitized signal(s)
227_40w3d.hea 2016-03-02 19:27 8.3K header file
229_39w1d.dat 2016-03-02 19:26 8.4M digitized signal(s)
229_39w1d.hea 2016-03-02 19:27 8.3K header file
230_38w5d.dat 2016-03-02 19:26 9.5M digitized signal(s)
230_38w5d.hea 2016-03-02 19:27 8.3K header file
232_37w6d.dat 2016-03-02 19:26 9.5M digitized signal(s)
232_37w6d.hea 2016-03-02 19:27 8.3K header file
233_39w3d.dat 2016-03-02 19:26 8.9M digitized signal(s)
233_39w3d.hea 2016-03-02 19:27 8.3K header file
234_37w5d.dat 2016-03-02 19:26 9.8M digitized signal(s)
234_37w5d.hea 2016-03-02 19:27 8.3K header file
235_39w0d.dat 2016-03-02 19:26 5.7M digitized signal(s)
235_39w0d.hea 2016-03-02 19:27 8.2K header file
237_40w2d.dat 2016-03-02 19:26 8.9M digitized signal(s)
237_40w2d.hea 2016-03-02 19:27 8.3K header file
DOI 2019-02-06 12:53 19
MD5SUMS 2016-03-02 19:51 2.6K
RECORDS 2016-03-02 19:36 266 list of record names
SHA1SUMS 2016-03-02 19:51 3.0K
SHA256SUMS 2016-03-02 19:51 4.3K
figure1.png 2016-03-03 18:29 54K
figure2.png 2016-03-03 18:29 64K
sensorpositions.dat 2016-03-02 19:26 894 digitized signal(s)
sensorpositions.hea 2016-03-02 19:27 9.0K header file
|
If you would like help understanding, using, or downloading content, please see our Frequently Asked Questions. If you have any comments, feedback, or particular questions regarding this page, please send them to the webmaster. Comments and issues can also be raised on PhysioNet's GitHub page. Updated Friday, 28 October 2016 at 16:58 EDT |
PhysioNet is supported by the National Institute of General Medical Sciences (NIGMS) and the National Institute of Biomedical Imaging and Bioengineering (NIBIB) under NIH grant number 2R01GM104987-09.
|