Basic Time and Frequency Domain Measures
Software and related material: Joseph E. Mietus, B.S.
Margret and H.A. Rey Institute for Nonlinear Dynamics in Physiology and Medicine
Division of Interdisciplinary Medicine and Biotechnology and Division of Cardiology
Beth Israel Deaconess Medical Center/Harvard Medical School, Boston, MA
I. Background
Heart rate variability (HRV) analysis attempts to assess cardiac autonomic regulation through quantification of sinus rhythm variability. The sinus rhythm times series is derived from the QRS to QRS (RR) interval sequence of the electrocardiogram (ECG), by extracting only normal sinus to normal sinus (NN) interbeat intervals. Relatively high frequency variations in sinus rhythm reflect parasympathetic (vagal) modulation, and slower variations reflect a combination of both parasympathetic and sympathetic modulation and non-autonomic factors [1-5].
Traditional heart rate variability (HRV) measures are usually divided into two broad categories: time domain measures and frequency domain measures [3,4]. The time domain heart rate variability statistics commonly calculated are defined in Table 1. Note, however, that computing pNNx with x < 50 ms in both long- and short-term recordings may provide a more robust index of fluctuations due to vagal tone than the standard pNN50 statistic [6]. Commonly used frequency domain measures are defined in Table 2.
The low frequency band (0.04 - 0.15 Hz) includes physiologic oscillations associated with baroreceptor reflexes and the high frequency band (0.15 - 0.40 Hz) encompasses respiratory sinus arrhythmia. The powers in these bands has been used to provide indexes of autonomic function. Such measures must be interpreted with caution, however. As noted, oscillations in the "low" frequency bands appear to be mediated by parasympathetic and sympathetic components, while the "high" frequency power is mediated exclusively by the vagus.
Traditionally, frequency domain measures are calculated by resampling the original NN interval series and then applying the fast Fourier transform or autoregressive spectral estimation (the maximum entropy method). This resampling, however, can cause an attenuation in the high frequency components. If discontinuities exist in the NN interval series, either because of the presence of abnormal beats or because of gaps or extreme noise in the original ECG recording, traditional approaches require either discarding the data or guesswork to estimate the locations of missing normal beats. To eliminate the need for evenly sampled data required by Fourier or maximum entropy methods, frequency domain spectra can be calculated using the Lomb periodogram for unevenly sampled data [7,8] (the method used in this toolkit).
Although the long term (24-hour) statistics of SDANN, SDNNIDX and ULF power can be calculated for shorter data lengths, they will become increasingly unreliable. For short-term data (less than 15 minutes in length), only the time domain measures of AVNN, SDNN, rMSSD and pNN50 and the frequency domain measures of total power, VLF power, HF power and LF/HF ratio should be used.
A number of the HRV measures are highly correlated with each other. These include SDNN, SDANN, total power and ULF power; SDNNIDX, VLF power and LF power; and rMSSD, pNN50 and HF power. The LF/HF ratio does not correlate strongly with any other HRV measures [4].
Heart rate variability (HRV) has been widely applied in basic and clinical research studies. Its clinical application is very limited at present, however. These limitations are due to lack of standardization of methodology and application to different non-comparable subsets of subjects, as well as to the confounding effects of age, gender, drugs, health status, and chronobiologic variations, among others. Furthermore, outliers due to ectopy and artifact can have major effects on computed HRV values. In elderly subjects, especially, a spuriously high value of certain measures may be due to the effects of "erratic supraventricular rhythm" [9] due to subtle atrial ectopy, wandering atrial pacemaker, or sinus node conduction abnormalities. Additional information on heart rate dynamics and analysis techniques, including non-linear and complexity based measures, can be found in the HRV 2006 course notes and elsewhere on PhysioNet (see for example: Detrended Fluctuation Analysis, Multiscale Entropy Analysis, and Information-Based Similarity, among others).
PhysioNet's HRV Toolkit, available here, is a rigorously validated package of open source software for HRV analysis, including visualization of NN interval time series, automated outlier removal, and calculation of the basic time- and frequency-domain HRV statistics widely used in the literature, including all of those listed in the tables below.
Several other high-quality, freely available HRV toolkits may also be of interest to researchers; links to them are provided at the end of this page.
Table 1: Commonly used time-domain measures
| AVNN* | Average of all NN intervals |
| SDNN* | Standard deviation of all NN intervals |
| SDANN | Standard deviation of the averages of NN intervals in all 5-minute segments of a 24-hour recording |
| SDNNIDX | Mean of the standard deviations of NN intervals in all 5-minute segments of a 24-hour recording |
| rMSSD* | Square root of the mean of the squares of differences between adjacent NN intervals |
| pNN50* | Percentage of differences between adjacent NN intervals that are greater than 50 ms; a member of the larger pNNx family [6] |
Table 2: Commonly used frequency-domain measures
| TOTPWR* | Total spectral power of all NN intervals up to 0.04 Hz |
| ULF | Total spectral power of all NN intervals up to 0.003 Hz |
| VLF* | Total spectral power of all NN intervals between 0.003 and 0.04 Hz |
| LF* | Total spectral power of all NN intervals between 0.04 and 0.15 Hz. |
| HF* | Total spectral power of all NN intervals between 0.15 and 0.4 Hz |
| LF/HF* | Ratio of low to high frequency power |
Selected References:
[1] Wolf MM, Varigos GA, Hunt D, Sloman JG. Sinus arrhythmia in acute myocardial infarction. Med J Aust 1978;2:52-53.
[2] Kleiger RE, Miller JP, Bigger JT, Moss AJ, and the Multicenter Post-Infarction Research Group. Decreased heart rate variability and its association with increased mortality after acute myocardial infarction. Am J Cardiol 1987;59:256-262.
[3] Task Force of the European Society of Cardiology and the North American Society of Pacing and Electrophysiology. Heart rate variability: Standards of measurement, physiological interpretation, and clinical use. Circulation 1996; 93:1043-1065.
[4] Mietus JE. Time domain measures: from variance to pNNx. http://physionet.org/events/hrv-2006/mietus-1.pdf
[5] Parati G, Mancia G, Di Rienzo M, Castiglioni P, Taylor JA, Studinger P. Point-Counterpoint: Cardiovascular variability is/is not an index of autonomic control of circulation. J Appl Physiol 2006; 101: 676 - 682.
[6] Mietus JE, Peng C-K, Henry I, Goldsmith RL, Goldberger AL. The pNNx-files: Reexamining a widely-used heart rate variability measure. Heart 2002;88:378-380.
[7] Press WH, Teukolsky SA, Vetterling WT, Flannery BP. Numerical Recipes in C: The Art of Scientific Computing, 2nd ed. Cambridge Univ. Press, 1992, pp. 575-584.
[8] Moody GB. Spectral analysis of heart rate without resampling. Computers in Cardiology 1993; 715-718.
[9] Stein PK, Yanez D, Domitrovich PP, Gottdiener J, Chaves P, Kronmal R, Rautaharju P. Heart rate variability is confounded by the presence of erratic sinus rhythm. Computers in Cardiology 2002; 669-72.
II. Obtaining the HRV Toolkit
The HRV Toolkit: Contents
The HRV Toolkit consists of a Usages file, a Makefile, and the following programs:
| plt_rrs | a shell script for plotting RR/NN interval series |
| get_hrv | a shell script for calculating time and frequency domain HRV statistics |
| Scripts above use the programs below, and others from the WFDB and plt packages | |
| rrlist.c | C code for extracting an RR interval list from an annotation file |
| filt.c | C code for filtering RR/NN intervals |
| filtnn.c | C code for filtering RR/NN intervals |
| statnn.c | C code for calculating time domain statistics |
| pwr.c | C code for calculating power in up to 10 frequency bands from a power spectrum |
| seconds.c | C code for converting hh:mm:ss to seconds |
| hours.c | C code for converting seconds to hh:mm:ss |
plt_rrs and get_hrv are the main scripts used in calculating HRV as illustrated below. These two scripts call the various C programs to accomplish their calculations. The C programs can be used separately and their usage and various options can be found in Usages, or by running any of these programs with the -h option.
Downloading and Installing the HRV Toolkit
Installing the HRV toolkit is easy:
- On Windows, install the free Cygwin software first; see our tutorial for details. Cygwin includes the gcc C compiler and all other utilities needed to build and run the components of the HRV Toolkit on Windows. Start a Cygwin (terminal emulator) window and use it for all of the remaining steps.
- Install the free WFDB and plt software packages.
- The HRV toolkit is available as a tarball of sources (for all platforms) or as tarballs of prebuilt binaries for GNU/Linux (x86), Mac OS X (x86), Solaris (SPARC), or Windows/Cygwin. Download the version of your choice.
- Unpack the HRV toolkit tarball you downloaded. (See the PhysioNet FAQ for information on unpacking tarballs.)
-
If you downloaded the sources, enter the source directory (HRV.src)
and compile and install the toolkit by typing:
make installIf you downloaded the binaries, move the contents of the HRV directory into some directory in your PATH (or add the HRV directory to your PATH). The binaries require the same additional packages as the source distribution (WFDB, plt, and (on Windows) Cygwin).
III. Using the HRV toolkit
User interface
The HRV toolkit does not include a graphical user interface. Its components are command-line tools that must be run from a terminal window (under MS-Windows, a Cygwin window) or by a shell script. (Even plt_rrs must be started from the command line or a script, although it produces graphical output.)Input data format
Both plt_rrs (for plotting the RR/NN interval series) and get_hrv (for calculating the HRV statistics) can take as input either a PhysioBank-compatible beat annotation file or a text file containing an RR interval list. RR interval lists can be in any of four formats:
- 3 columns (T, RR, A)
- 2 columns (RR, A)
- 2 columns (T, RR)
- 1 column (RR)
Although T is assumed to be expressed in seconds by default, the -I option of plt_rrs and get_hrv (see below) can be used if the RR interval list contains T values expressed as clock time (hh:mm:ss.xxx), hours (hh.xxxxxxx) or minutes (mm.xxxxx). Similarly, although RR is assumed to be expressed in seconds, intervals in milliseconds can be input by using the `-m' option. (See details below or in Usages).
Beat annotation files are available for most of the PhysioBank records that include ECGs. For information about record and annotation conventions see the PhysioNet FAQ. If you wish to study a recording for which no beat annotation file or RR interval listing is available, you may be able to create an annotation file using software from the WFDB software package. Additional information on RR intervals, heart rate, and HRV can be found at the RR/HR/HRV Howto.
Input used in this tutorial
The examples below read the ecg annotations from record chf03 of the BIDMC Congestive Heart Failure Database. To reproduce the results shown below, it is not necessary to download this annotation file, because applications that use the WFDB library (including those in the HRV Toolkit that read annotation files) can locate and read the copy from the PhysioNet web server if no local copy exists. In order to do so successfully, it is necessary to specify the path to the record from the PhysioBank archive directory within the record name, as shown in these examples (use chfdb/chf03 rather than simply chf03). Examples that illustrate input from RR interval lists assume that the input file is named chf03.rr, and that it is in the current (local) directory.
plt_rrs: plotting the RR/NN interval time series
NN interval outliers due to missed or false beat detections can seriously corrupt HRV statistics. Most frequency domain measures are especially susceptible to outliers, particularly LF and HF power which can be in error by greater than 1000%. Most time domain measures are less affected by outliers, but still can give errors in excess of 100%. AVNN, pNN50 and ULF power are least affected, generally with errors less than 10% (see Time domain Measures: From Variance to pNNx).
To reduce the errors due to outliers, the first step in HRV analysis is to visually examine the RR/NN interval data and determine the necessity of outlier filtering. This can be done using plt_rrs.
plt_rrs can be used with either an annotation file or an RR interval listing (see below) and has the following options:
plt_rrs [options] -R rrfile | record annotator [start [end]]
Plot RR intervals or RR interval heart rates
options :
[-P 2|4|8|16|24|32] : plot 2, 4, 8, 16, 24 or 32 hours per page
(default: scale page length to data length)
[-R rrfile] : RR interval file : time (sec), interval
[-N] : plot NN intervals instead of RR intervals
[-H] : plot RR/NN interval heart rate
[-F "filt hwin"] : filter intervals, plot filtered data
[-f "filt hwin"] : filter intervals
[-p] : plot points
[-I c|h|m] : input time format: hh::mm:ss, hours, minutes (default: seconds)
[-m] : RR intervals in msec
[-y "ymin ymax"] : y axis limits
[-o] : output postscript
By default, plt_rrs will open a plt window and
display the plot on the terminal screen. To output the plot as a
postscript file use the -o option. To print the plot to a
postscript printer use
plt_rrs -o other arguments | lpr
Using an annotation file, like this:
plt_rrs chfdb/chf03 ecg
will plot the entire RR interval series for the congestive heart failure record
chfdb/chf03 using the ecg
annotator. By default, plt_rrs scales the plot so that the
entire RR interval series is plotted on one page, with a maximum of 8
lines per page. To plot a specified number of hours per page, use the
-P option. For example, -P 4 will print 4 hours per page (15
minutes per line), -P 16 will print 16 hours per page (2 hours per
line), etc.
To plot selected portions of the data, specify a start time and optionally an end time. For example,
plt_rrs -P 8 chfdb/chf03 ecg 00:00:00 01:00:00
plots the first hour of the RR interval sequence:
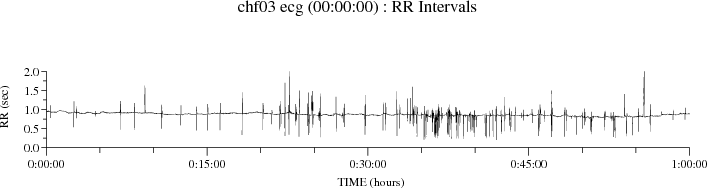
With the -p option, as in
plt_rrs -P 8 -p chfdb/chf03 ecg 00:00:00 01:00:00
plt_rrs plots the RR interval sequence as individual points:
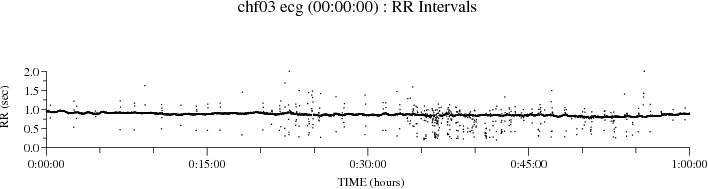
With the -N option, as in
plt_rrs -P 8 -N chfdb/chf03 ecg 00:00:00 01:00:00
plt_rrs plots NN intervals only, omitting intervals that are
not bounded by normal sinus beats at both ends:
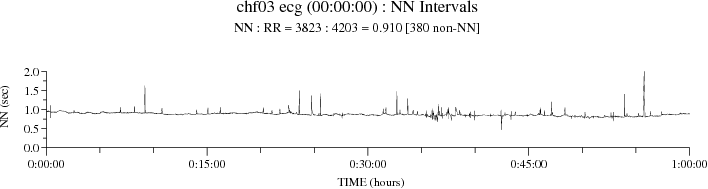
The "NN : RR = 3823 : 4203 = 0.910 [380 non-NN]" in the title indicates the total number of NN intervals, the total number of RR intervals, the fraction of RR intervals that are NN intervals and the number of non-NN intervals.
To filter outliers, use the -F or -f options, as in
plt_rrs -P 8 -N -F "0.2 20 -x 0.4 2.0" chfdb/chf03 ecg 00:00:00 01:00:00
This procedure will plot the first hour of the filtered NN intervals
sequence. Here the "-F 0.2 20 -x 0.4 2.0" specifies the filtering
parameters as follows. First, any intervals less than 0.4 sec or
greater than 2.0 sec are excluded. Next, using a window of 41
intervals (20 intervals on either side of the central point), the
average over the window is calculated excluding the central
interval. If the central interval lies outside 20% (0.2) of the window
average this interval is flagged as an outlier and excluded. Then the
window is advanced to the next interval. These parameters can be
adjusted as appropriate for different data sets.
Using -F allows plt_rrs to plot the excluded intervals as small filled circles:
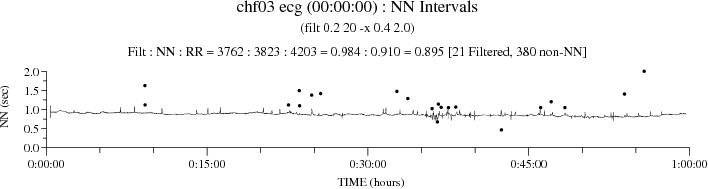
The -f option suppresses output of excluded intervals:

The "Filt : NN : RR = 3762 : 3823 : 4203 = 984 : 0.910 = 0.895 [61 Filtered, 380 non-NN]" in the title of the two plots above gives the total number of NN intervals remaining after filtering, the total number of NN intervals before filtering, the total number of RR intervals, the fraction of NN intervals remaining after filtering, the fraction of RR intervals that are NN intervals, the fraction of the total number of RR intervals that are NN intervals remaining after filtering, and the number of NN intervals filtered out together with the number of non-NN intervals.
Filtering the NN interval data may not be necessary if there are no extreme outliers.
To plot the RR interval series of an RR interval file containing three columns of data : time in seconds, interval in seconds and beat label, use the -R option followed by the name of the file, rather than specifying the record and annotator names. For example, if the file chf03.rr contains:
1.356 0.952 N 2.312 0.956 N 3.252 0.940 N 4.212 0.960 N 5.156 0.944 N 6.112 0.956 N 7.048 0.936 N 7.996 0.948 N 8.960 0.964 N 9.900 0.940 N . . .use plt_rrs -R chf03.rr with any of the above options to plot its contents.
The above RR interval list was generated using the command
rrlist ecg chfdb/chf03 -s >chf03.rr
(Notice that the record and annotator arguments appear in reverse
order in rrlist commands.) The various other options
available for rrlist are given in Usages.
If an RR interval listing contains only time in seconds and interval in seconds, e.g.,
1.356 0.952 2.312 0.956 3.252 0.940 4.212 0.960 5.156 0.944 6.112 0.956 7.048 0.936 7.996 0.948 8.960 0.964 9.900 0.940 . . .then a command such as plt_rrs -R chf03.rr plots the RR interval sequence correctly, but will not be able to plot the NN interval sequence using the -N option, since the beat labels are missing.
To plot an RR interval listing containing only RR intervals in seconds and beat labels, e.g.,
0.952 N 0.956 N 0.940 N 0.960 N 0.944 N 0.956 N 0.936 N 0.948 N 0.964 N 0.940 N . . .the command plt_rrs -R chf03.rr calculates the time of each RR interval from the RR interval sequence, making the assumption that the intervals are consecutive.
Finally, if the RR interval listing contains only RR intervals in seconds, e.g.,
0.952 0.956 0.940 0.960 0.944 0.956 0.936 0.948 0.964 0.940 . . .then plt_rrs -R chf03.rr calculates the time of the time of the RR interval from the RR interval sequence, but as in a previous example will not be able to plot the NN interval sequence using the -N option.
If the RR intervals in any of the above formats are listed in milliseconds
rather than seconds, use plt_rrs with the -m option.
get_hrv: calculating the HRV statistics
To calculate the time and frequency domain HRV statistics use get_hrv, where the options are:
get_hrv [options] -R rrfile | record annotator [start [end]]
Get HRV statistics :
REC : NN/RR AVNN SDNN SDANN SDNNINDX RMSSD PNN : TOTPWR ULF VLF LF HF LF/HF
options :
[-R rrfile] : RR interval file : time (sec), interval
[-f "filt hwin"] : filter outliers
[-p "nndiff ..."] : nn difference for pnn (default: 50 msec)
[-P "lo1 hi1 lo2 hi2 lo3 hi3 lo4 hi4"] : power bands
(default : 0 0.0033 0.0033 0.04 0.04 0.15 0.15 0.4)
[-s] : short term stats of
REC : NN/RR AVNN SDNN RMSSD PNN : TOTPWR VLF LF HF LF/HF
[-I c|h|m] : input time format: hh::mm:ss, hours, minutes (default: seconds)
[-m] : RR intervals in msec
[-M] : output statistics in msec rather than sec
[-L] : output statistics on one line
[-S] : plot HRV results on screen
plotting options :
[-F "filt hwin"] : filter outliers, plot filtered data
[-y "ymin ymax"] : time series y-axis limits ("- -" for self-scaling)
[-X maxfreq] : fft maximum frequency (default : 0.4 Hz)
[-Y fftmax] : fft maximum ("-" for self-scaling)
[-o] : output plot in postscript
get_hrv uses statnn to calculate the time domain statistics,
and lomb (from the WFDB software package) and pwr to calculate
the frequency domain statistics. NN/RR is the fraction of total RR intervals
that are classified as normal-to-normal (NN) intervals and included in the
calculation of HRV statistics. This ratio can be used as a measure of data
reliability. For example, if the NN/RR ratio is less than 0.8, fewer than 80%
of the RR intervals are classified as NN intervals, and the results will be
somewhat unreliable.
The command
get_hrv -f "0.2 20 -x 0.4 2.0" -p "20 50" chfdb/chf03 ecg
should give the following output:
chfdb/chf03 :
NN/RR = 0.942899
AVNN = 0.892206
SDNN = 0.054712
SDANN = 0.0466773
SDNNIDX = 0.0241124
rMSSD = 0.0177694
pNN20 = 0.0688698
pNN50 = 0.0252389
TOT PWR = 0.0033882
ULF PWR = 0.00270489
VLF PWR = 0.00039018
LF PWR = 0.000124104
HF PWR = 0.000169027
LF/HF = 0.734226
In this case AVNN, SDNN, SDANN, SDNNIDX are rMSSD are given in seconds,
pNN values as ratios and PWR values in seconds squared.
Note that the exact values obtained on different platforms may vary by
approximately 10-6 due to round-off errors.
By default, get_hrv outputs the HRV statistics in seconds. To output results in milliseconds use the -M option, as in
get_hrv -M -f "0.2 20 -x 0.4 2.0" -p "20 50" chfdb/chf03 ecg
This should give the following output:
chfdb/chf03 :
NN/RR = 0.942899
AVNN = 892.206
SDNN = 54.712
SDANN = 46.6773
SDNNIDX = 24.1124
rMSSD = 17.7694
pNN20 = 6.88698
pNN50 = 2.52389
TOT PWR = 3388.65
ULF PWR = 2705.19
VLF PWR = 390.25
LF PWR = 124.156
HF PWR = 169.057
LF/HF = 0.734403
In this case AVNN, SDNN, SDANN, SDNNIDX are rMSSD are given in milliseconds,
pNN values as percentages and PWR values in milliseconds squared.
To output the HRV statistics all on one line, use the -L option:
get_hrv -L -M -f "0.2 20 -x 0.4 2.0" -p "20 50" chfdb/chf03 ecg
This will output
chfdb/chf03 : 0.942899 892.206 54.712 46.6773 24.1124 17.7694 6.88698 2.52389 : 3388.65 2705.19 390.25 124.156 169.057 0.734403
where the measures are
REC : NN/RR AVNN SDNN SDANN SDNNINDX RMSSD PNN20 PNN50 : TOTPWR ULF VLF LF HF LF/HF
To limit the output to the statistics named as short-term HRV statistics in tables 1 and 2 above, use the -s option, as in
get_hrv -s -M -f "0.2 20 -x 0.4 2.0" -p "20 50" chfdb/chf03 ecg 0:00:00 1:00:00
This should give the following output:
chfdb/chf03 :
NN/RR = 0.90173
AVNN = 867.958
SDNN = 39.7191
rMSSD = 20.8659
pNN20 = 6.26553
pNN50 = 2.20811
TOT PWR = 1702.96
VLF PWR = 1423.01
LF PWR = 103.941
HF PWR = 176.008
LF/HF = 0.590547
To output the short-term HRV statistics all on one line, use both the -s and the -L options:
get_hrv -L -s -M -f "0.2 20 -x 0.4 2.0" -p "20 50" chfdb/chf03 ecg 0:00:00 1:00:00
This will output
chfdb/chf03 : 0.90173 867.958 39.7191 20.8659 6.26553 2.20811 : 1702.96 1423.01 103.941 176.008 0.590547
where the measures are
REC : NN/RR AVNN SDNN RMSSD PNN20 PNN50 : TOTPWR VLF LF HF LF/HF
As with plt_rrs, get_hrv with the -R option can accept a text file containing an RR interval listing in any of the formats described above. If the interval times are not listed, get_hrv will calculate the time of the RR interval from the RR interval sequence, and if there are no beat labels, get_hrv -R will treat all intervals as normal.
If the RR intervals are listed in milliseconds rather than seconds, use
get_hrv with the -m option.
To plot the HRV results in a window including the NN interval time series, the NN interval histogram and the NN interval power spectrum use the -S option to get_hrv. As with plt_rrs the -o will output PostScript, and the -F filtering option will plot the excluded intervals of the NN interval time series: the non-normal beats as small open circles and outliers as small filled circles. For example
get_hrv -S -s -M -F "0.2 20 -x 0.4 2.0" -p "20 50" chf03 ecg 0:00:00 1:00:00
will generate the following plot:

Note that plots made this way show the calculated HRV statistics with reduced precision.
Additional get_hrv plotting options include -y for scaling the time series y-axis limits, the -X option for scaling the power spectrum x-axis limit and the -Y option for scaling the power spectrum y-axis limit.
IV. Representative measurements of HRV
Table 3 gives representative values of HRV measurements, obtained using the HRV toolkit from the 72 subjects in the MIT-BIH Normal Sinus Rhythm Database and the Normal Sinus Rhythm RR Interval Database. These representative values (comparable to those previously reported in healthy subjects) are not, however, intended as a standard normative database.
Table 3:
HRV in 24-hour NN interval time series from 72 ostensibly healthy subjects
(35 males, 37 females, ages 20-76 years, mean age 55)
| Measurement | Mean ± SD |
|---|---|
| AVNN (msec) | 787.7 ± 79.2 |
| SDNN (msec) | 136.5 ± 33.4 |
| SDANN (msec) | 127.2 ± 35.7 |
| SDNNIDX (msec) | 51.2 ± 14.2 |
| rMSSD (msec) | 27.9 ± 12.3 |
| pNN20 (%) | 34.2 ± 13.7 |
| pNN50 (%) | 7.5 ± 7.6 |
| TOTPWR (msec2) | 21490 ± 11577 |
| ULF PWR (msec2) | 18128 ± 10109 |
| VLF PWR (msec2) | 1900 ± 1056 |
| LF PWR (msec2) | 961 ± 722 |
| HF PWR (msec2) | 501 ± 847 |
| LF/HF ratio | 2.7 ± 1.3 |
HRV measurements obtained for each of these 72 subjects are available here.
Links to other HRV toolkits
Several other free HRV toolkits exist. Among them are:
- Physionet Cardiovascular Signal Toolbox (https://physionet.org/physiotools/physionet-cardiovascular-signal-toolbox/)
An open-source modular program for calculating heart rate variability (HRV) implemented in Matlab with evidence-based algorithms and output formats. The Toolbox is compatible with 64-bit MATLAB on GNU/Linux, Mac OS X, and MS-Windows. It was compared to several other open source and proprietary tools including the PhysioNet C HRV Toolkit to create a benchmark for the field. It was shown to be equivalent to the PhysioNet C HRV Toolkit. It contains the most extensive set of tools in any HRV algorithm collection so far published. It has no dependencies outside of Matlab (tested on Matlab R2017a and R2017b).
- KUBIOS-HRV (http://kubios.uku.fi/)
Authors: JP Niskanen and colleagues at Kuopio University, Finland
KUBIOS-HRV 2.0 is available for Windows and Linux. It is a standalone Matlab application (only the Matlab runtime compiler and libraries, both included in the package, are needed in order to use it). KUBIOS-HRV includes a well-written 53-page user guide with detailed descriptions of all of the algorithms used, mathematical derivations where needed, and an extensive bibliography.
The package includes automated and interactive methods for outlier identification and removal; a smoothness-based detrending algorithm for preprocessing time series; derivation of SDNN, SDSD, RMSSD, pNN50, HRV triangular index, and TINN; FFT and AR power spectrum estimates, VLF, LF, HF, and LF/HF ratio; and Poincaré plots, ApEn, SampEn, DFA, correlation dimension, and recurrence plots. Inputs are RR interval lists in text format.
KUBIOS-HRV 2.0 and its user guide were released in 2002, and it was described in detail in:
Juha-Pekka Niskanen, Mika P Tarvainen, Perttu O Ranta-aho, Pasi A Karjalainen, "Software for advanced HRV analysis." Computer Methods and Programs in Biomedicine 76(1):73-81 (2004 Oct; DOI: 10.1016/j.cmpb.2004.03.004)
- R-HRV (http://cran.r-project.org/web/packages/RHRV/)
Authors: M Lado, A Mendez, L Rodriguez-Linares, X Vila (Univ. of Vigo, Spain)
R-HRV is freely available in source form (for all platforms) and as prebuilt binaries for Mac OS X and Windows, together with a 13-page user guide. It requires R (a very widely used open-source statistics package). R-HRV includes functions for importing PhysioBank (WFDB) annotation files, automated outlier handling, and frequency domain HRV analysis.
R-HRV was described in:
L. Rodriguez-Linares, X. Vila, A. Mendez, M. Lado, D. Olivieri, "RHRV: An R-based software package for heart rate variability analysis of ECG recordings." 3rd Iberian Conference in Systems and Information Technologies (CISTI 2008), 21-23 June 2008.
- Software for Heart Rate Variability
(http://www.macalester.edu/~kaplan/hrv/doc/)
Authors: Danny Kaplan and Phil Staffin (Macalester College)
This software, written as Matlab m-code, includes routines for outlier handling, SDNN, RMSSD, a generalization of pNN50, spectral analysis using FFT (VLF, LF, HF, LF/HF), ApEn, and DFA. Inputs are RR interval lists in text format.